The Python Toolbox for Neurophysiological Signal Processing
This package is the continuation of NeuroKit 1. It's a user-friendly package providing easy access to advanced biosignal processing routines. Researchers and clinicians without extensive knowledge of programming or biomedical signal processing can analyze physiological data with only two lines of code.
import neurokit2 as nk
# Download example data
data = nk.data("bio_eventrelated_100hz")
# Preprocess the data (filter, find peaks, etc.)
processed_data, info = nk.bio_process(ecg=data["ECG"], rsp=data["RSP"], eda=data["EDA"], sampling_rate=100)
# Compute relevant features
results = nk.bio_analyze(processed_data, sampling_rate=100)And boom 💥 your analysis is done 😎
To install NeuroKit2, run this command in your terminal:
pip install neurokit2
If you're not sure how/what to do, be sure to read our installation guide.


NeuroKit2 is a collaborative project with a community of contributors with all levels of development expertise. Thus, if you have some ideas for improvement, new features, or just want to learn Python and do something useful at the same time, do not hesitate and check out the following guides:




Click on the links above and check out our tutorials:
- Get familiar with Python in 10 minutes
- Recording good quality signals
- What software for physiological signal processing
- Install Python and NeuroKit
- Included datasets
- Additional Resources
- Simulate Artificial Physiological Signals
- Customize your Processing Pipeline
- Event-related Analysis
- Interval-related Analysis
- Analyze Electrodermal Activity (EDA)
- Analyze Respiratory Rate Variability (RRV)
- Extract and Visualize Individual Heartbeats
- Locate P, Q, S and T waves in ECG
- Complexity Analysis of Physiological Signals
- Analyze Electrooculography EOG data
- Fit a function to a signal
You can try out these examples directly in your browser.
Don't know which tutorial is suited for your case? Follow this flowchart:
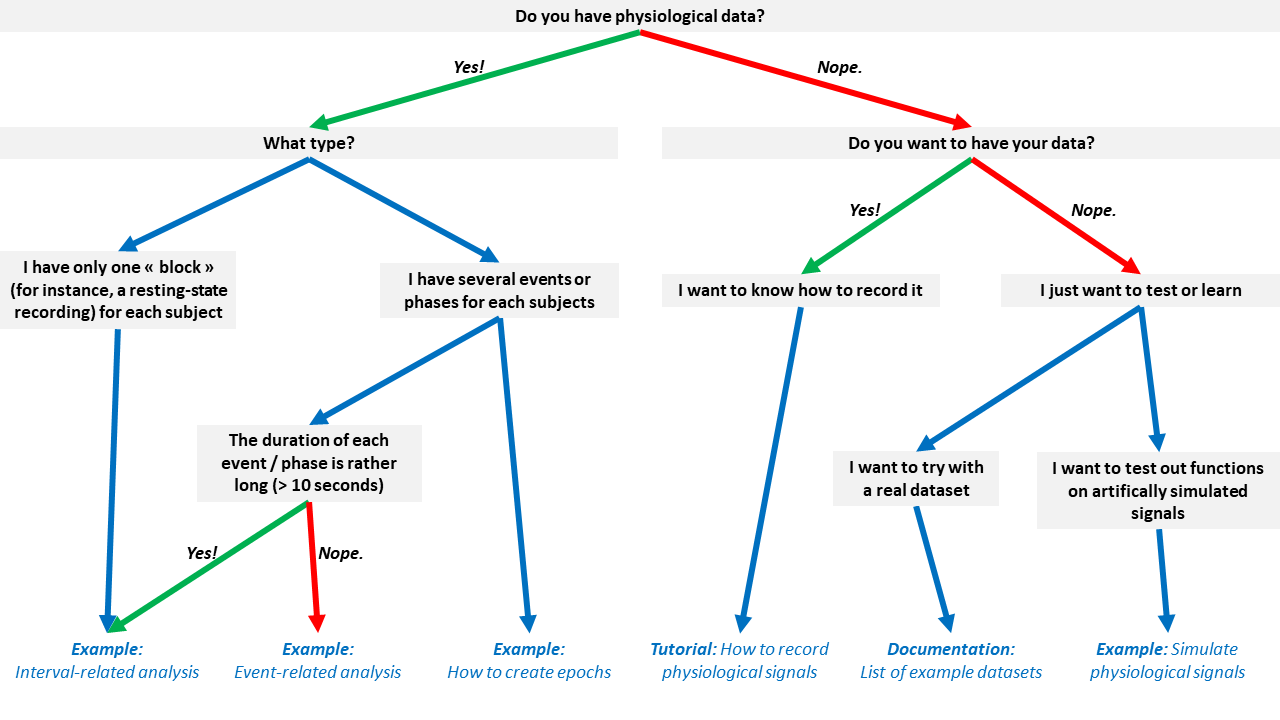

nk.cite()You can cite NeuroKit2 as follows:
- Makowski, D., Pham, T., Lau, Z. J., Brammer, J. C., Lesspinasse, F., Pham, H.,
Schölzel, C., & S H Chen, A. (2020). NeuroKit2: A Python Toolbox for Neurophysiological
Signal Processing. Retrieved March 28, 2020, from https://github.com/neuropsychology/NeuroKit
Full bibtex reference:
@misc{neurokit2,
doi = {10.5281/ZENODO.3597887},
url = {https://github.com/neuropsychology/NeuroKit},
author = {Makowski, Dominique and Pham, Tam and Lau, Zen J. and Brammer, Jan C. and Lespinasse, Fran\c{c}ois and Pham, Hung and Schölzel, Christopher and S H Chen, Annabel},
title = {NeuroKit2: A Python Toolbox for Neurophysiological Signal Processing},
publisher = {Zenodo},
year = {2020},
}import numpy as np
import pandas as pd
import neurokit2 as nk
# Generate synthetic signals
ecg = nk.ecg_simulate(duration=10, heart_rate=70)
ppg = nk.ppg_simulate(duration=10, heart_rate=70)
rsp = nk.rsp_simulate(duration=10, respiratory_rate=15)
eda = nk.eda_simulate(duration=10, scr_number=3)
emg = nk.emg_simulate(duration=10, burst_number=2)
# Visualise biosignals
data = pd.DataFrame({"ECG": ecg,
"PPG": ppg,
"RSP": rsp,
"EDA": eda,
"EMG": emg})
nk.signal_plot(data, subplots=True)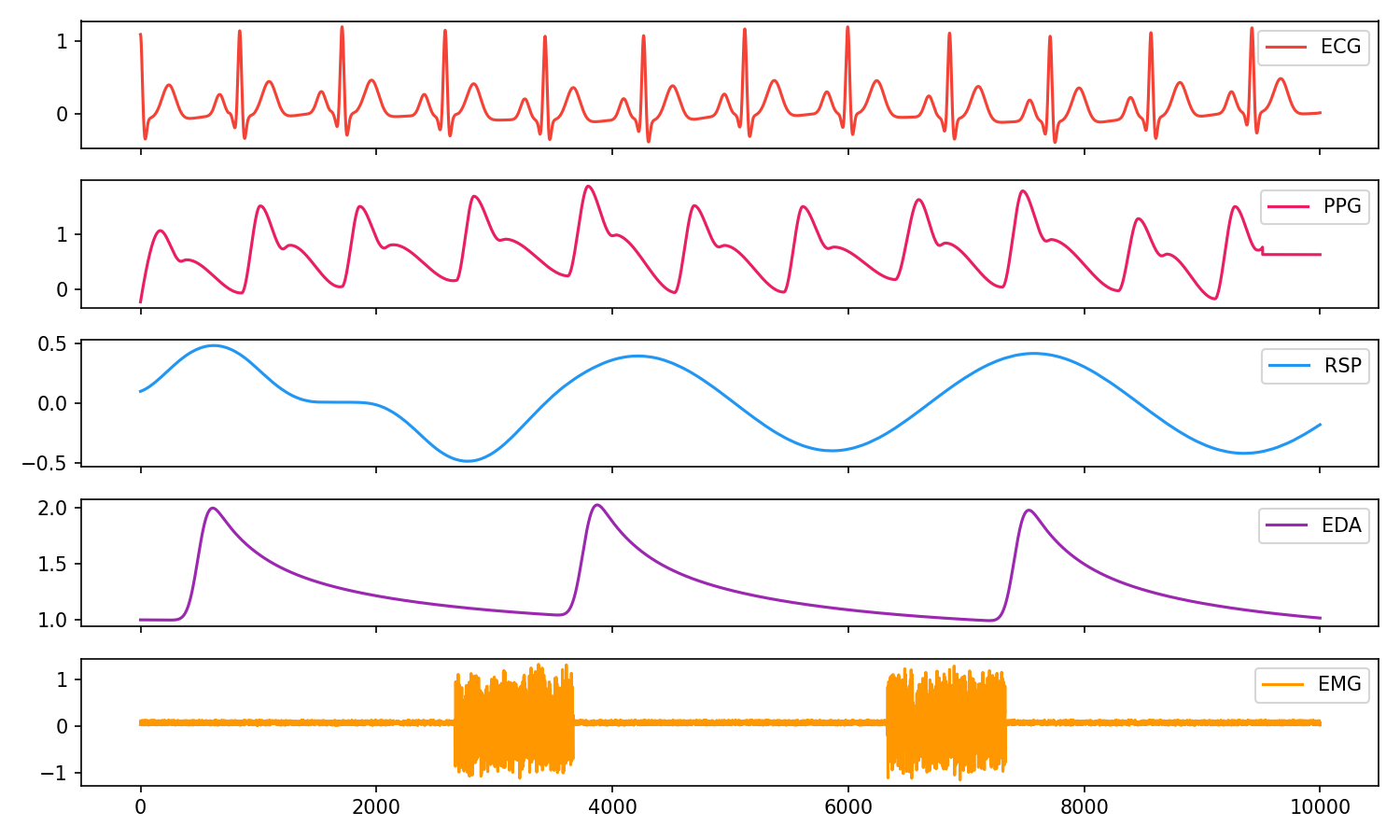
# Generate 10 seconds of EDA signal (recorded at 250 samples / second) with 2 SCR peaks
eda = nk.eda_simulate(duration=10, sampling_rate=250, scr_number=2, drift=0.01)
# Process it
signals, info = nk.eda_process(eda, sampling_rate=250)
# Visualise the processing
nk.eda_plot(signals, sampling_rate=250)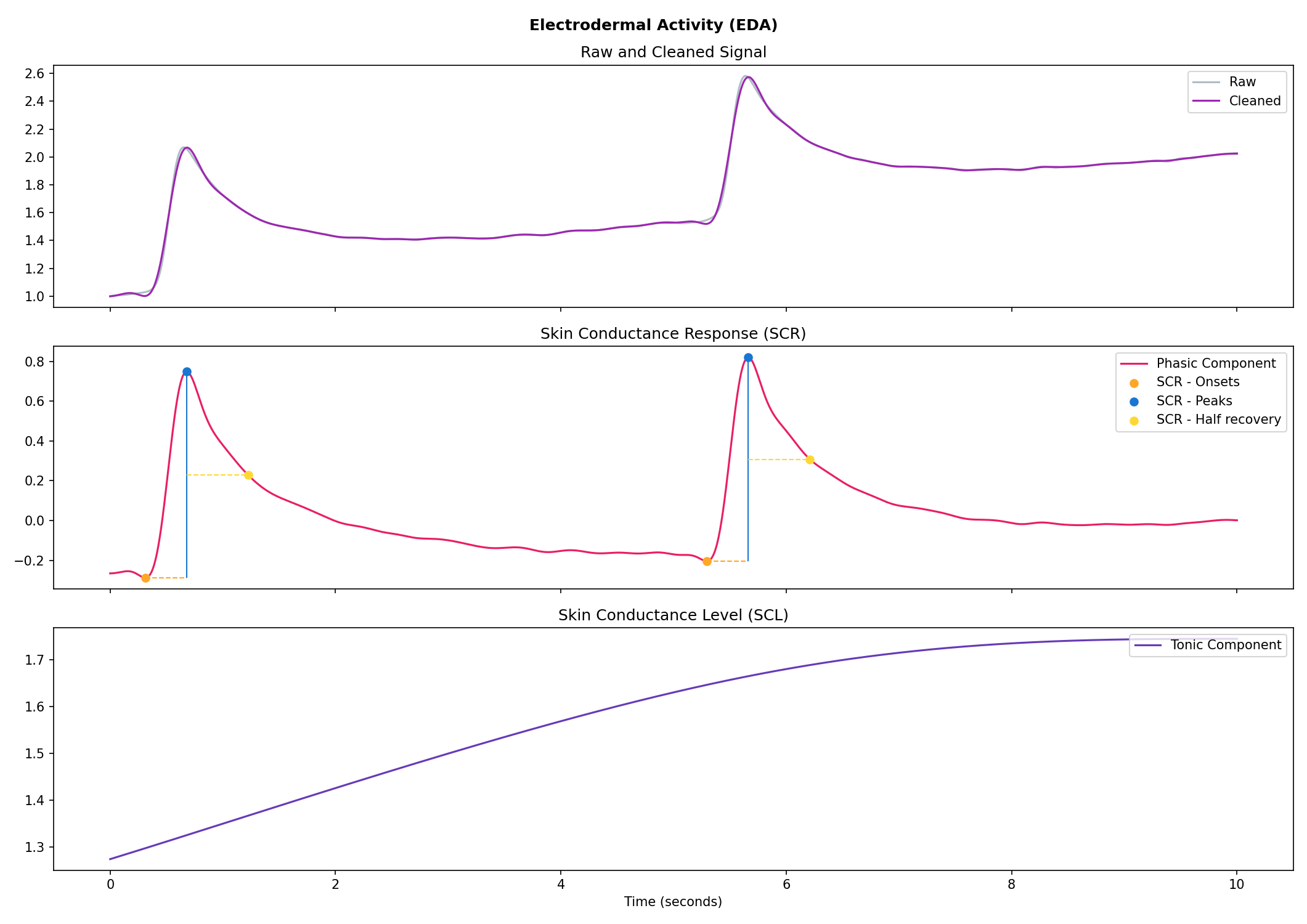
# Generate 15 seconds of ECG signal (recorded at 250 samples / second)
ecg = nk.ecg_simulate(duration=15, sampling_rate=250, heart_rate=70)
# Process it
signals, info = nk.ecg_process(ecg, sampling_rate=250)
# Visualise the processing
nk.ecg_plot(signals, sampling_rate=250)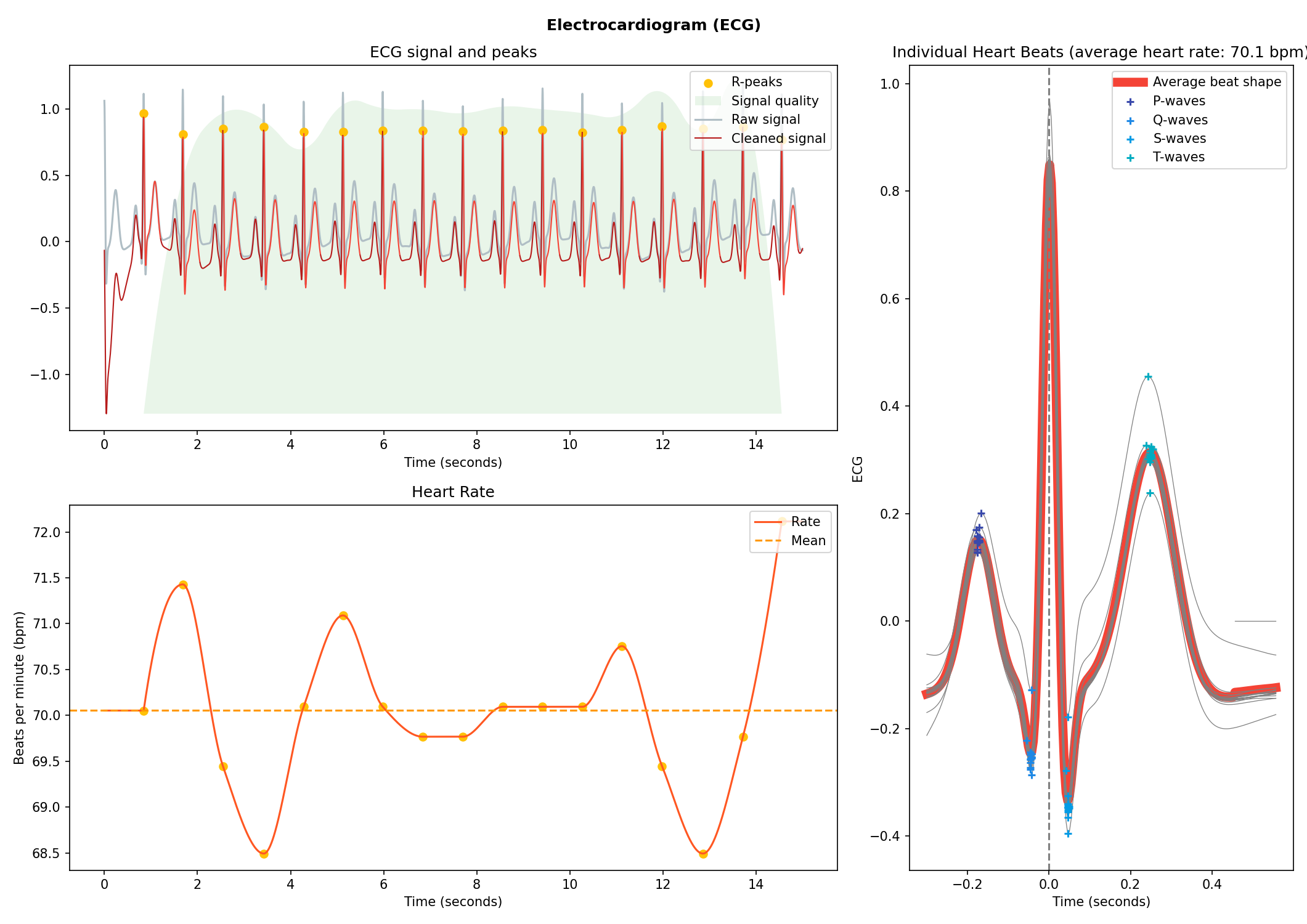
# Generate one minute of respiratory (RSP) signal (recorded at 250 samples / second)
rsp = nk.rsp_simulate(duration=60, sampling_rate=250, respiratory_rate=15)
# Process it
signals, info = nk.rsp_process(rsp, sampling_rate=250)
# Visualise the processing
nk.rsp_plot(signals, sampling_rate=250)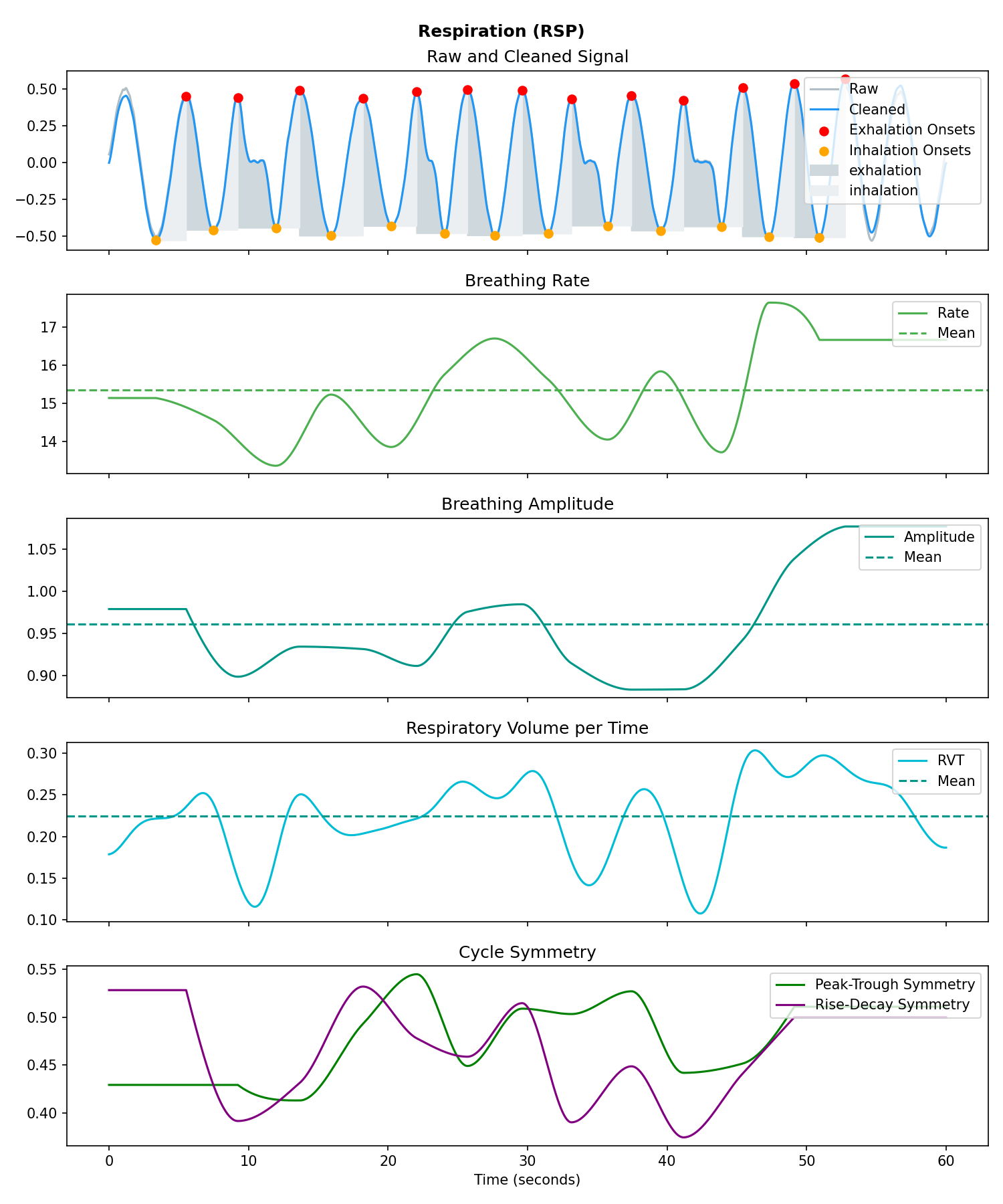
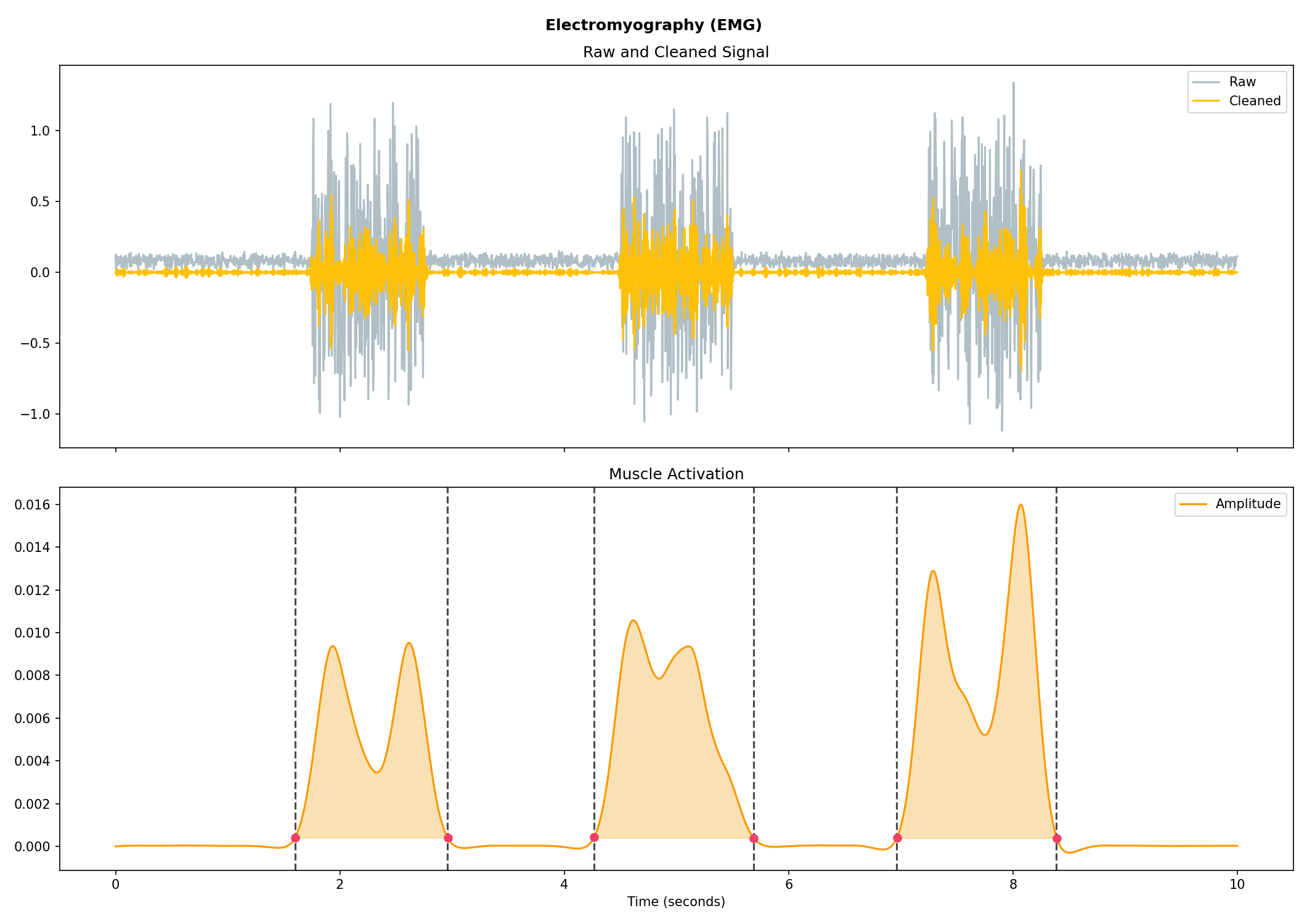
# Generate 10 seconds of EMG signal (recorded at 250 samples / second)
emg = nk.emg_simulate(duration=10, sampling_rate=250, burst_number=3)
# Process it
signal, info = nk.emg_process(emg, sampling_rate=250)
# Visualise the processing
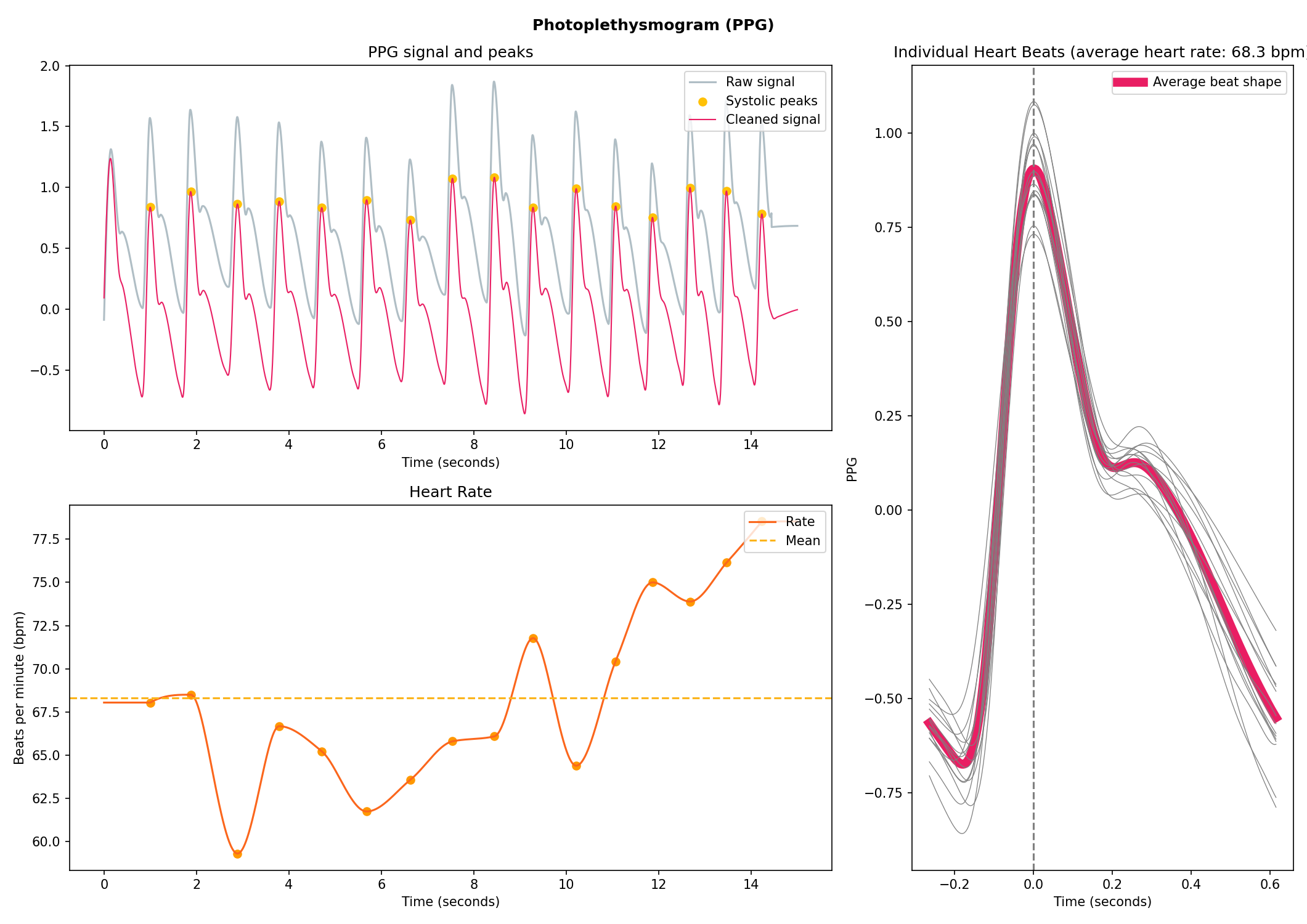
nk.emg_plot(signals, sampling_rate=250)# Generate 15 seconds of PPG signal (recorded at 250 samples / second)
ppg = nk.ppg_simulate(duration=15, sampling_rate=250, heart_rate=70)
# Process it
signals, info = nk.ppg_process(ppg, sampling_rate=250)
# Visualize the processing
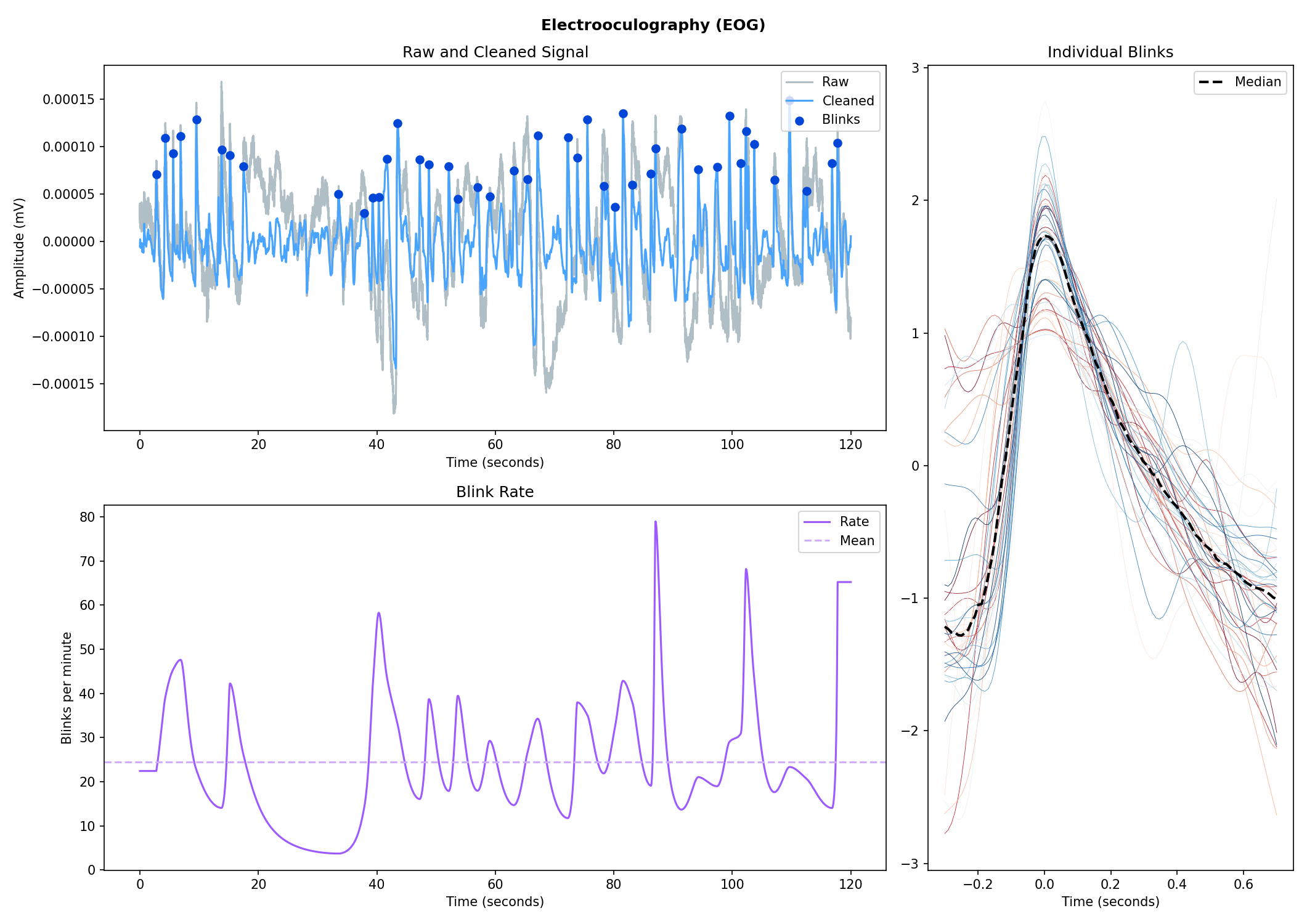
nk.ppg_plot(signals, sampling_rate=250)# Import EOG data
eog_signal = nk.data("eog_100hz")
# Process it
signals, info = nk.eog_process(eog_signal, sampling_rate=100)
# Plot
plot = nk.eog_plot(signals, sampling_rate=100)Consider helping us develop it!
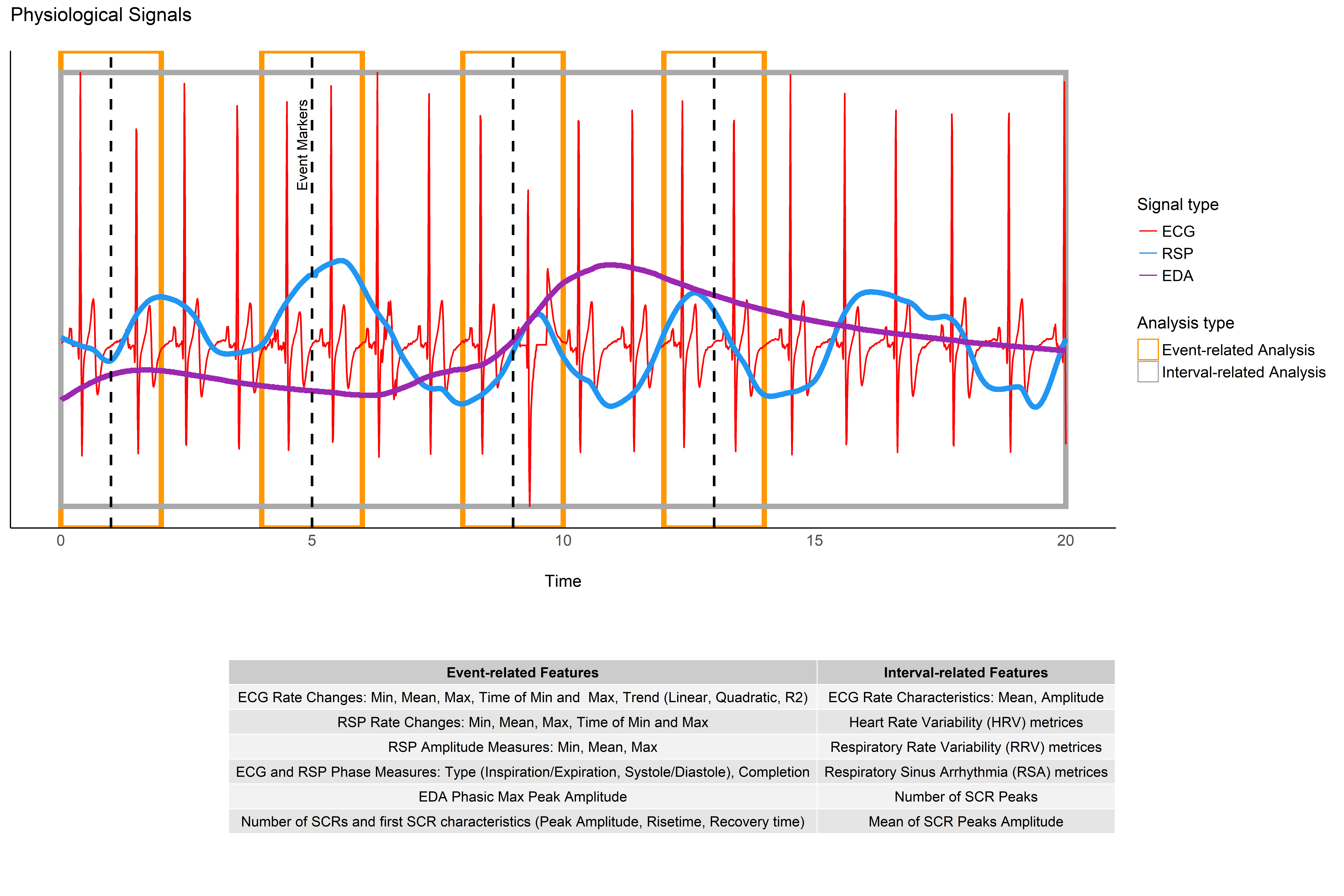
The analysis of physiological data usually comes in two types, event-related or interval-related.
This type of analysis refers to physiological changes immediately occurring in response to an event. For instance, physiological changes following the presentation of a stimulus (e.g., an emotional stimulus) indicated by the dotted lines in the figure above. In this situation the analysis is epoch-based. An epoch is a short chunk of the physiological signal (usually < 10 seconds), that is locked to a specific stimulus and hence the physiological signals of interest are time-segmented accordingly. This is represented by the orange boxes in the figure above. In this case, using bio_analyze() will compute features like rate changes, peak characteristics and phase characteristics.
This type of analysis refers to the physiological characteristics and features that occur over longer periods of time (from a few seconds to days of activity). Typical use cases are either periods of resting-state, in which the activity is recorded for several minutes while the participant is at rest, or during different conditions in which there is no specific time-locked event (e.g., watching movies, listening to music, engaging in physical activity, etc.). For instance, this type of analysis is used when people want to compare the physiological activity under different intensities of physical exercise, different types of movies, or different intensities of stress. To compare event-related and interval-related analysis, we can refer to the example figure above. For example, a participant might be watching a 20s-long short film where particular stimuli of interest in the movie appears at certain time points (marked by the dotted lines). While event-related analysis pertains to the segments of signals within the orange boxes (to understand the physiological changes pertaining to the appearance of stimuli), interval-related analysis can be applied on the entire 20s duration to investigate how physiology fluctuates in general. In this case, using bio_analyze() will compute features such as rate characteristics (in particular, variability metrices) and peak characteristics.
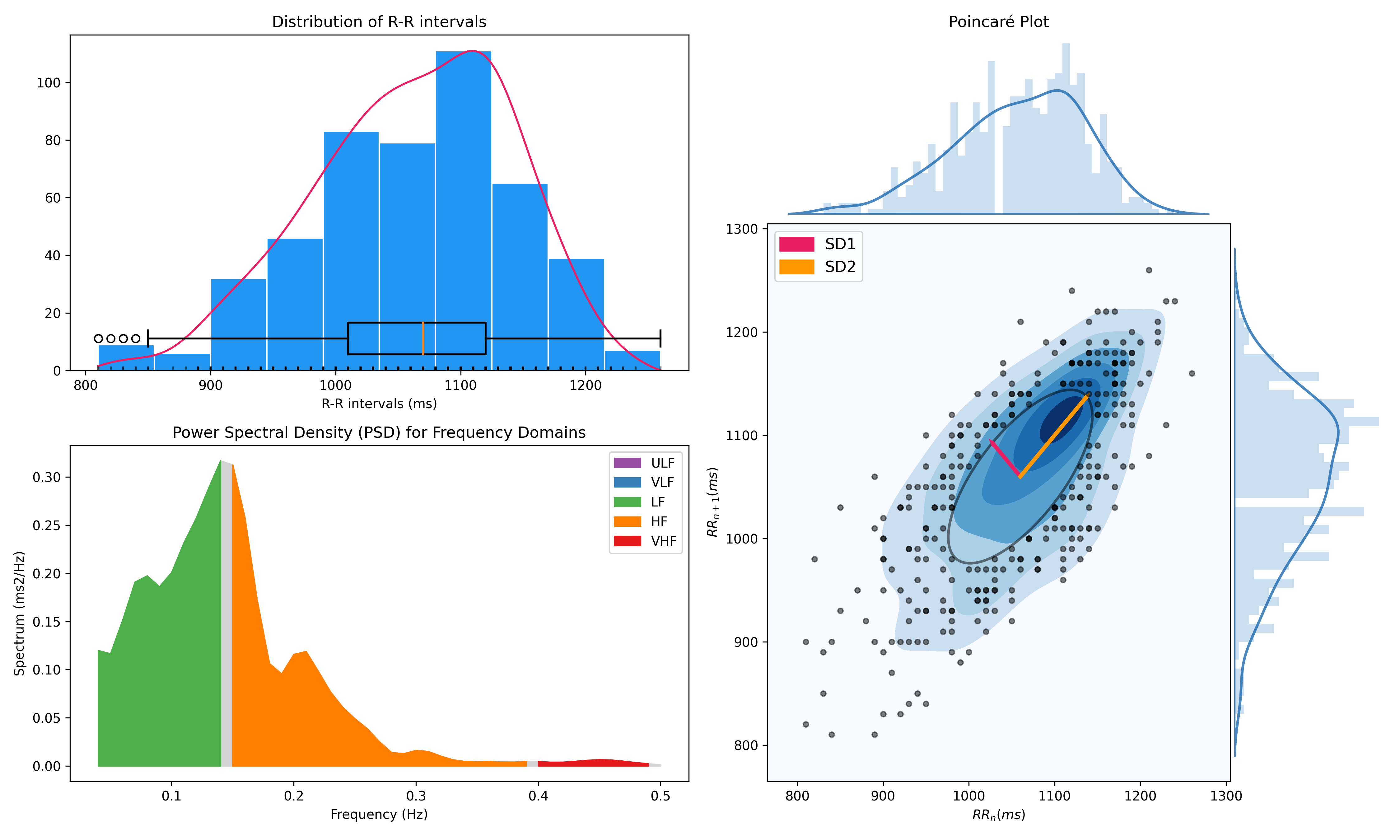
- Compute HRV indices
- Time domain: RMSSD, MeanNN, SDNN, SDSD, CVNN etc.
- Frequency domain: Spectral power density in various frequency bands (Ultra low/ULF, Very low/VLF, Low/LF, High/HF, Very high/VHF), Ratio of LF to HF power, Normalized LF (LFn) and HF (HFn), Log transformed HF (LnHF).
- Nonlinear domain: Spread of RR intervals (SD1, SD2, ratio between SD2 to SD1), Cardiac Sympathetic Index (CSI), Cardial Vagal Index (CVI), Modified CSI, Sample Entropy (SampEn).
# Download data
data = nk.data("bio_resting_8min_100hz")
# Find peaks
peaks, info = nk.ecg_peaks(data["ECG"], sampling_rate=100)
# Compute HRV indices
nk.hrv(peaks, sampling_rate=100, show=True)
>>> HRV_RMSSD HRV_MeanNN HRV_SDNN ... HRV_CVI HRV_CSI_Modified HRV_SampEn
>>> 0 69.697983 696.395349 62.135891 ... 4.829101 592.095372 1.259931- Delineate the QRS complex of an electrocardiac signal (ECG) including P-peaks, T-peaks, as well as their onsets and offsets.
# Download data
ecg_signal = nk.data(dataset="ecg_3000hz")['ECG']
# Extract R-peaks locations
_, rpeaks = nk.ecg_peaks(ecg_signal, sampling_rate=3000)
# Delineate
signal, waves = nk.ecg_delineate(ecg_signal, rpeaks, sampling_rate=3000, method="dwt", show=True, show_type='all')
- Signal processing functionalities
- Filtering: Using different methods.
- Detrending: Remove the baseline drift or trend.
- Distorting: Add noise and artifacts.
# Generate original signal
original = nk.signal_simulate(duration=6, frequency=1)
# Distort the signal (add noise, linear trend, artifacts etc.)
distorted = nk.signal_distort(original,
noise_amplitude=0.1,
noise_frequency=[5, 10, 20],
powerline_amplitude=0.05,
artifacts_amplitude=0.3,
artifacts_number=3,
linear_drift=0.5)
# Clean (filter and detrend)
cleaned = nk.signal_detrend(distorted)
cleaned = nk.signal_filter(cleaned, lowcut=0.5, highcut=1.5)
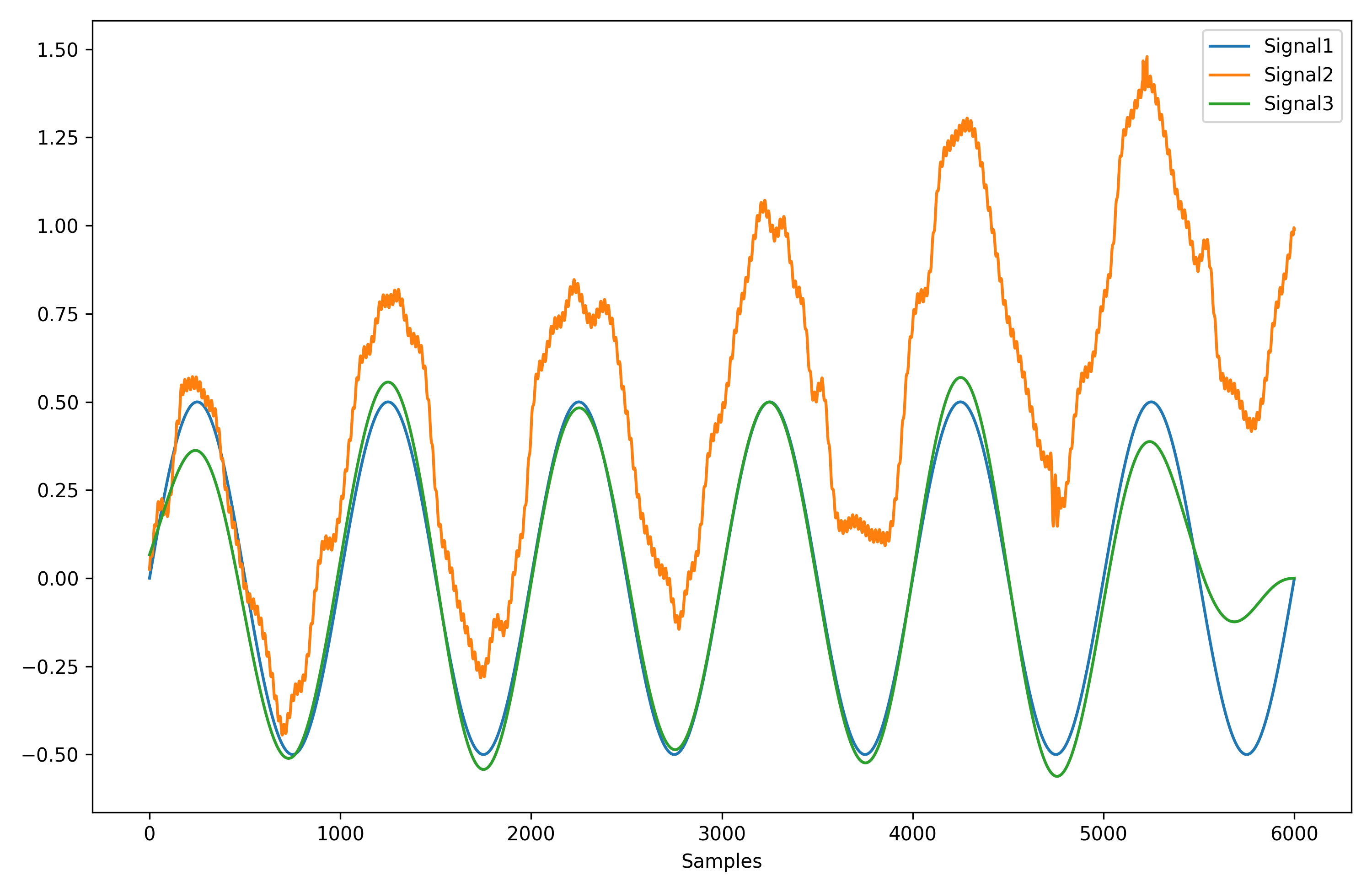
# Compare the 3 signals
plot = nk.signal_plot([original, distorted, cleaned])- Optimize complexity parameters (delay tau, dimension m, tolerance r)
# Generate signal
signal = nk.signal_simulate(frequency=[1, 3], noise=0.01, sampling_rate=100)
# Find optimal time delay, embedding dimension and r
parameters = nk.complexity_optimize(signal, show=True)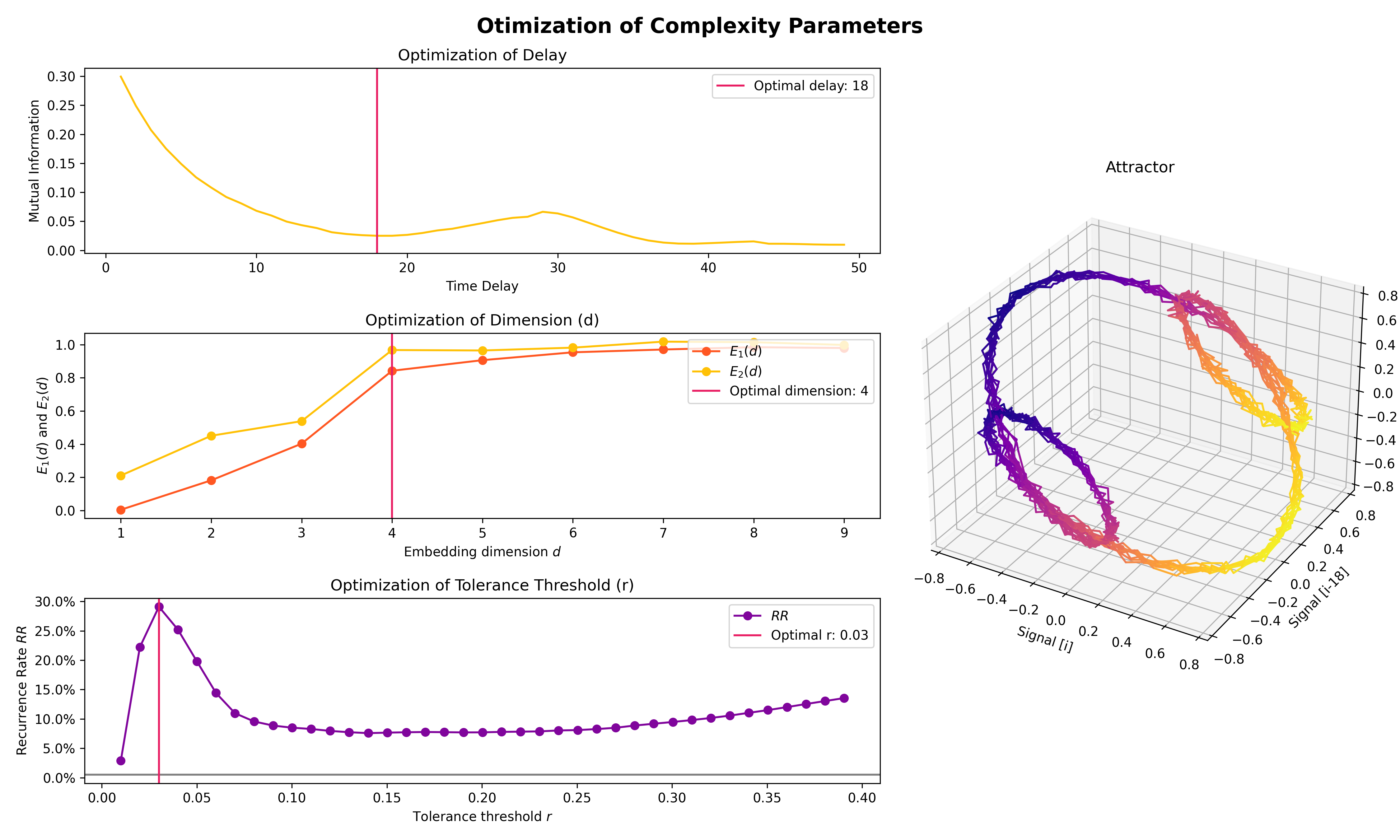
- Compute complexity features
- Entropy: Sample Entropy (SampEn), Approximate Entropy (ApEn), Fuzzy Entropy (FuzzEn), Multiscale Entropy (MSE), Shannon Entropy (ShEn)
- Fractal dimensions: Correlation Dimension D2, ...
- Detrended Fluctuation Analysis
nk.entropy_sample(signal)
nk.entropy_approximate(signal)# Create complex signal
signal = nk.signal_simulate(duration=10, frequency=1) # High freq
signal += 3 * nk.signal_simulate(duration=10, frequency=3) # Higher freq
signal += 3 * np.linspace(0, 2, len(signal)) # Add baseline and linear trend
signal += 2 * nk.signal_simulate(duration=10, frequency=0.1, noise=0) # Non-linear trend
signal += np.random.normal(0, 0.02, len(signal)) # Add noise
# Decompose signal using Empirical Mode Decomposition (EMD)
components = nk.signal_decompose(signal, method='emd')
nk.signal_plot(components) # Visualize components
# Recompose merging correlated components
recomposed = nk.signal_recompose(components, threshold=0.99)
nk.signal_plot(recomposed) # Visualize components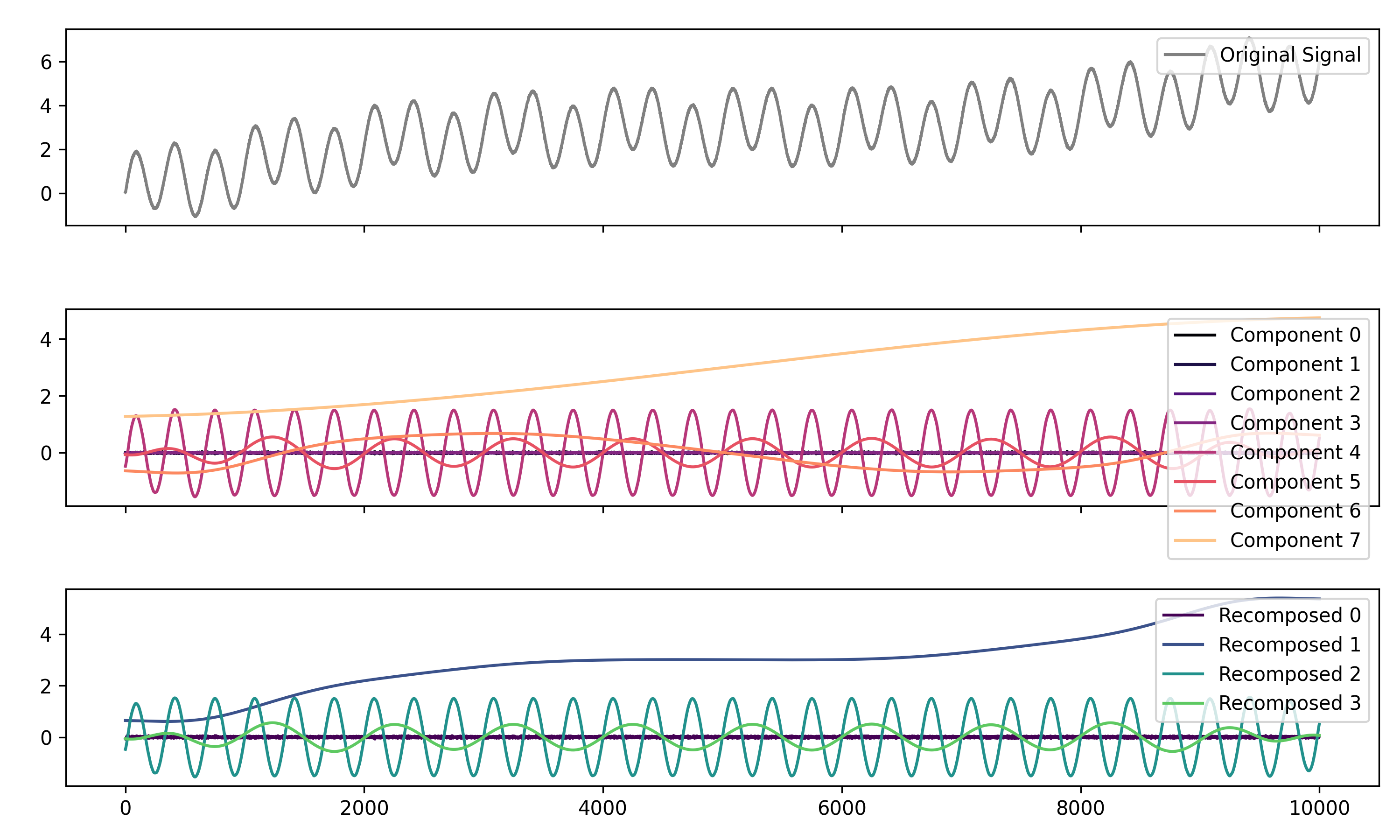
# Generate signal with frequencies of 5, 20 and 30
signal = nk.signal_simulate(frequency=5) + 0.5*nk.signal_simulate(frequency=20) + nk.signal_simulate(frequency=30)
# Find Power Spectrum Density with different methods
# Mutlitaper
multitaper = nk.signal_psd(signal, method="multitapers", show=False, max_frequency=100)
# Welch
welch = nk.signal_psd(signal, method="welch", min_frequency=1, show=False, max_frequency=100)
# Burg
burg = nk.signal_psd(signal, method="burg", min_frequency=1, show=False, ar_order=15, max_frequency=100)
# Visualize the different methods together
fig, ax = plt.subplots()
ax.plot(welch["Frequency"], welch["Power"], label="Welch", color="#CFD8DC", linewidth=2)
ax.plot(multitaper["Frequency"], multitaper["Power"], label="Multitaper", color="#00695C", linewidth=2)
ax.plot(burg["Frequency"], burg["Power"], label="Burg", color="#0097AC", linewidth=2)
ax.set_title("Power Spectrum Density (PSD)")
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)")
ax.set_ylabel("PSD (ms^2/Hz)")
ax.legend(loc="upper right")
# Plot 3 frequencies of generated signal
ax.axvline(5, color="#689F38", linewidth=3, ymax=0.95, linestyle="--")
ax.axvline(20, color="#689F38", linewidth=3, ymax=0.95, linestyle="--")
ax.axvline(30, color="#689F38", linewidth=3, ymax=0.95, linestyle="--")
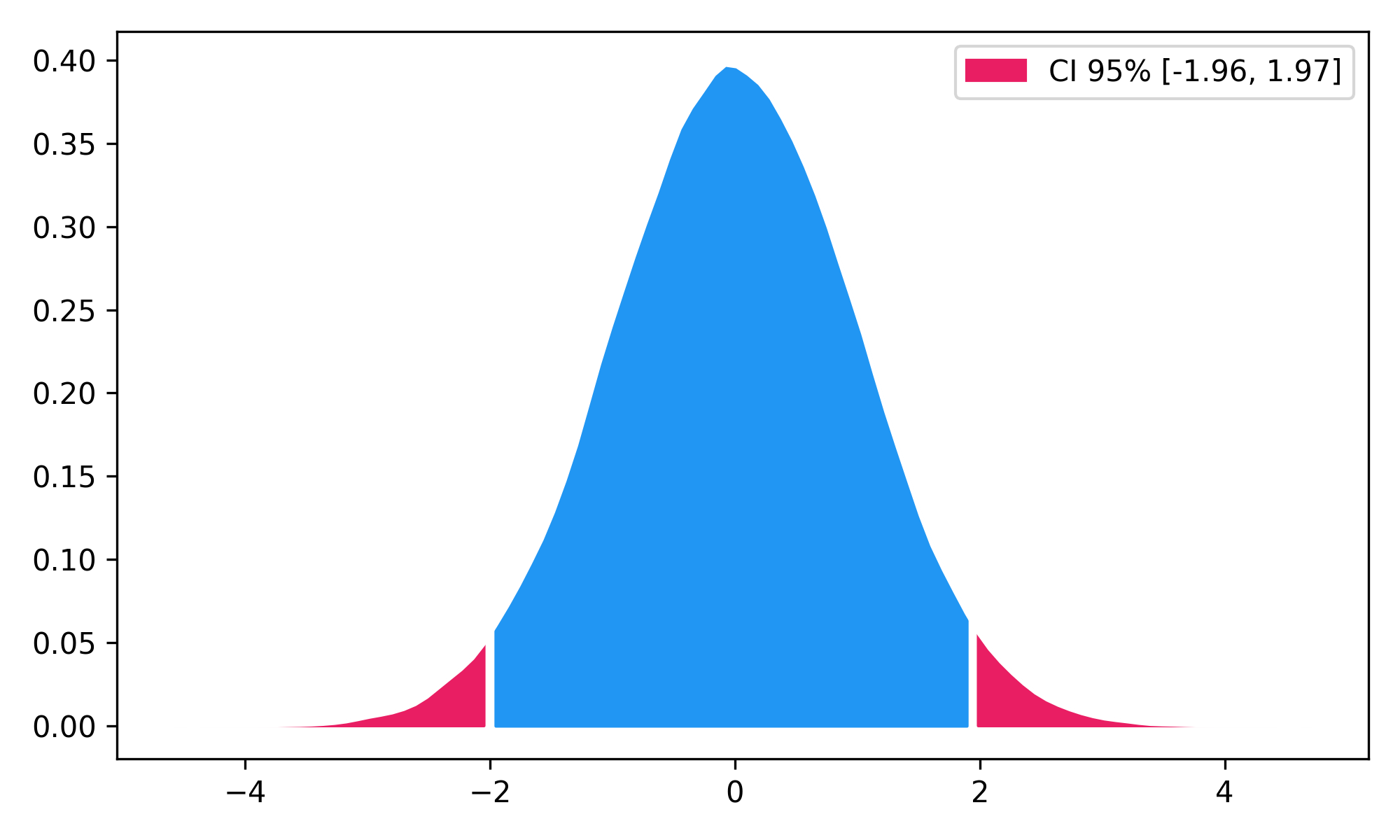
- Highest Density Interval (HDI)
x = np.random.normal(loc=0, scale=1, size=100000)
ci_min, ci_max = nk.hdi(x, ci=0.95, show=True)


The authors do not provide any warranty. If this software causes your keyboard to blow up, your brain to liquify, your toilet to clog or a zombie plague to break loose, the authors CANNOT IN ANY WAY be held responsible.